
Neuropharmacology
-
Professor
Department of Pharmacology and ToxicologyPhone: (409) 772-6292
Fax: (409) 772-6293
Email: garudenk@utmb.edu
Lab Webpage -
University of Groningen, Netherlands (PhD)
UT Southwestern Medical Center/HHMI (Post-doctoral Studies) -
There are an estimated hundred billion neurons in the human brain and they are connected to each other via physical contact points called synapses. Synapses enable neurons to communicate with each other. The hundreds of trillions of synapses in our brain establish neural circuitries that guide how we think, move and feel. More than a thousand different proteins are found at synapses and they form complex protein networks. Paradoxically, synapses are both insoluble and yet also plastic. On the one hand, synapses are isolated biochemically as the 'triton-insoluble' fraction. Yet on the other hand, in vivo, synapses come and go. Synapses grow 'weaker' and 'stronger', as their adhesive properties and their ability to transmit signals change. Significantly, properties of synapses also appear to change as a function of their activity. External stimuli such as events triggering memory and learning, stress, and exposure to chemicals such as drugs of abuse, anti-depressants and anti-psychotics, all seem to affect synapses and the connections they form. Many different neuropsychiatric disorders and neurodegenerative disorders are increasingly being referred to as 'synaptopathies', emphasizing the role of disrupted synaptic structure and function in the pathogenesis of these disorders. By unraveling how the many different synaptic proteins interact with each other and form complex protein networks, we hope to not only gain fundamental insight into how neurons communicate with each other enabling the brain to function, but also to discover new potential therapeutic targets.
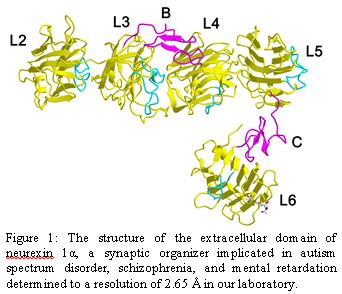
Our laboratory is particularly fascinated by the complex protein networks in the synaptic cleft found at chemical synapses, i.e. the 250 space between the 'pre-synaptic' membrane which hosts the exocytosis machinery for synaptic vesicles and the 'post-synaptic' membrane which hosts machinery responding to the transmitted chemical signals. We are studying a number of synaptic adhesion molecules and synaptic organizers to understand their role in mediating synapse formation, maintenance, and plasticity. One family of synaptic adhesion molecules that we have studied extensively is the family of neurexins. Neurexins play a role in synapse organization and adhesion. Mutations and lesions in neurexins have recently been implicated in autism spectrum disorder, schizophrenia and mental retardation (Fig. 1).
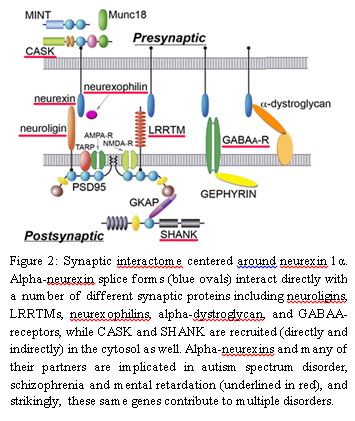
Excitingly, not only neurexins, but also many of their direct protein partners in the synaptic cleft are implicated in these diseases as well (Fig. 2). Neurexins and their partners must touch fundamental biological processes that are involved in the pathogenesis of these disorders, but it is not clear which processes these are and the exact role that neurexins and their partners play in these processes.
Our laboratory is working to understand on a molecular level how neurexins, their partners, as well as a number of other synaptic organizers recognize, bind, and arrange different synaptic partners in the synaptic cleft impacting synaptic function. By understanding the molecular mechanisms of these molecules, we will be able to not only further delineate their role at synapses but also understand why these molecules, when disrupted, contribute to neurological disorders. We use biochemical and biophysical techniques as well as protein crystallography.
-
39. Molecular mechanism of contactin 2 homophilic interaction. In Structure. Elsevier BV. Fan, S., Liu, J., Chofflet, N., Bailey, A. O., Russell, W. K., Zhang, Z., Takahashi, H., Ren, G., & Rudenko, G. (2024).
38. Chofflet N, Naito Y, Pastore AJ, et al. Structural and functional characterization of the IgSF21-neurexin2α complex and its related signaling pathways in the regulation of inhibitory synapse organization. Front Mol Neurosci. 2024;17:1371145. Published 2024 Mar 20.
37. Designer molecules of the synaptic organizer MDGA1 reveal 3D conformational control of biological function. Lee H, Chofflet N, Liu J, Fan S, Lu Z, Resua Rojas M, Penndorf P, Bailey AO, Russell WK, Machius M, Ren G, Takahashi H, Rudenko G. J Biol Chem. 2023, Apr;299(4):104586.
Selected as JBC's "Editors' Picks as a paper providing an exceptional contribution to the field
36. Kumar A, Aglyamova G, Yim YY, et al. Chemically targeting the redox switch in AP1 transcription factor ΔFOSB. Nucleic Acids Res. 2022;50(16):9548-9567.
35. Interplay between hevin, SPARC, and MDGAs: modulators of neurexin-neuroligin transsynaptic bridges. Fan S, Gangwar SP, Machius M, Rudenko G. Structure. 2021 Jul 1;29(7):664-678.e6. doi: 10.1016/j.str.2021.01.003.
Highlight: Juan J. Ramirez, Dhanesh Sivadasan Bindu, Cagla Eroglu. Building and destroying synaptic bridges: How do Hevin/Sparcl1, SPARC, and MDGAs modify trans-synaptic neurexin-neuroligin interactions? Structure, 2021, Jul 1; 29(7):635-637.34. Highly Conserved Molecular Features in IgLONs Contrast Their Distinct Structural and Biological Outcomes. Venkannagari H, Kasper JM, Misra A, Rush SA, Fan S, Lee H, Sun H, Seshadrinathan S, Machius M, Hommel JD, Rudenko G. J Mol Biol. 432(19):5287-5303 (2020) doi: 10.1016/j.jmb.2020.07.014.
33. Discovery of phenanthridine analogues as novel chemical probes disrupting the binding of DNA to ΔFosB homodimers and ΔFosB/JunD heterodimers. Li Y, Liu Z, Aglyamova G, Chen J, Chen H, Bhandari M, White MA, Rudenko G, Zhou J. Bioorg Med Chem Lett. 30(16):127300 (2020). doi: 10.1016/j.bmcl.2020.127300.
32. Self-assembly of the bZIP transcription factor ΔFosB. Yin Z, Venkannagaria H, Lynch H, Aglyamova G, Bhandari M, Machius M, Nestler EJ, Robison AJ, Rudenko G. Current Research in Structural Biology. 2:1-13 (2020)
31. Editorial overview: Synapses: bridging the gap between their structure and function. Rudenko G, Takahashi, H. Curr Opin Struct Biol.54:iii-vii. (2019)
30. Neurexins - versatile molecular platforms in the synaptic cleft. Rudenko G. Curr Opin Struct Biol. 54:112-121 (2019)
29. Structural Plasticity of Neurexin 1α: Implications for its Role as Synaptic Organizer. Liu J, Misra A, Reddy MVVVS, White MA, Ren G, Rudenko G. J Mol Biol. 430(21):4325-4343 (2018)
28. Activator Protein-1: redox switch controlling structure and DNA-binding. Yin Z, Machius M, Nestler EJ, and Rudenko G. Nucleic Acids Research Sept. 7 (2017)
27. Molecular Mechanism of MDGA1: Regulation of Neuroligin 2:Neurexin Trans-synaptic Bridges. Gangwar SP, Zhong X, Seshadrinathan S, Chen H, Machius M, and Rudenko G. Neuron Jun. 21 (2017)
26. Molecular and Cellular Mechanisms of Synaptopathies. Ardiles AO, Grabrucker AM, Scholl FG, Rudenko G, and Borsello T. Neural Plasticity Apr. 30 (2017)
25. Dynamic Control of Synaptic Adhesion and Organizing Molecules in Synaptic Plasticity. Rudenko G. Neural Plasticity Jan. 31 (2017)
24. Molecular Architecture of Contactin-associated Protein-like 2 (CNTNAP2) and its Interaction with Contactin 2 (CNTN2). Lu Z, Reddy MV, Liu J, Kalichava A, Liu J, Zhang L, Chen F, Wang Y, Holthauzen LM, White MA, Seshadrinathan S, Zhong X, Ren G, and Rudenko G. The Journal of Biological Chemistry Sept. 12 (2016)
23. Functional analysis of rare variants found in schizophrenia implicates a critical role for GIT1-PAK3 signaling in neuroplasticity. Kim MJ, Biag J, Fass DM, Lewis MC, Zhang Q, Fleishman M, Gangwar SP, Machius M, Fromer M, Purcell SM, McCarroll SA, Rudenko G, Premont RT, Scolnick EM, and Haggarty SJ. Mol Psychiatry July 26 (2016)
22. Data publication with the structural biology data grid supports live analysis. Meyer PA, Socias S, Key J, Ransey E, Tjon EC, Buschiazzo A, Lei M, Botka C, Withrow J, Neau D, Rajashankar K, Anderson KS, Baxter RH, Blacklow SC, Boggon TJ, Bonvin AM, Borek D, Brett TJ, Caflisch A, Chang CI, Chazin WJ, Corbett KD, Cosgrove MS, Crosson S, Dhe-Paganon S, Di Cera E, Drennan CL, Eck MJ, Eichman BF, Fan QR, Ferré-D'Amaré AR, Christopher Fromme J, Garcia KC, Gaudet R, Gong P, Harrison SC, Heldwein EE, Jia Z, Keenan RJ, Kruse AC, Kvansakul M, McLellan JS, Modis Y, Nam Y, Otwinowski Z, Pai EF, Pereira PJ, Petosa C, Raman CS, Rapoport TA, Roll-Mecak A, Rosen MK, Rudenko G, Schlessinger J, Schwartz TU, Shamoo Y, Sondermann H, Tao YJ, Tolia NH, Tsodikov OV, Westover KD, Wu H, Foster I, Fraser JS, Maia FR, Gonen T, Kirchhausen T, Diederichs K, Crosas M, and Sliz P. Nat Commun. Mar. 07 (2016)
21. Calsyntenin3: Molecular Architecture and Interaction with Neurexin 1alpha. Lu Z, Wang Y, Chen F, Tong H, Reddy MV, Luo L, Seshadrinathan S, Zhang L, Holthauzen LM, Craig AM, Ren G, and Rudenko G. The Journal of Biological Chemistry Oct. 28 (2014)
20. Threonine 149 Phosphorylation Enhances ΔFosB Transcriptional Activity to Control Psychomotor Responses to Cocaine.Cates HM, Thibault M, Pfau M, Heller E, Eagle A, Gajewski P, Bagot R, Colangelo C, Abbott T, Rudenko G, Neve R, Nestler EJ, and Robison AJ. The Journal of Neuroscience 34(34):11461-11469 (2014)
19. The specific α-neurexin interactor calsyntenin-3 promotes excitatory and inhibitory synapse development. Pettem KL, Yokomaku D, Luo L, Linhoff MW, Prasad T, Connor SA, Siddiqui TJ, Kawabe H, Chen F, Zhang L, Rudenko G, Wang YT, Brose N, Craig AM. Neuron 80(1):113-28 (2013).
Highlight: Featured article by Neuron18. Small molecule screening identifies regulators of the transcription factor ΔFosB. Wang Y, Cesena TI, Ohnishi Y, Burger-Caplan R, Lam V, Kirchhoff PD, Larsen SD, Larsen MJ, Nestler EJ, Rudenko G. ACS Chem Neurosci. 3(7):546-56 (2012)
17. The structure of neurexin 1α reveals features promoting a role as synaptic organizer. Chen F, Venugopal V, Murray B, Rudenko G. Structure 19(6):779-89 (2011).
Highlight: Early Immediate Publication, Featured Article and Cover Highlight by Structure
Highlight: Comment in Structure 19(6):749-75016. Model of human low-density lipoprotein and bound receptor based on cryoEM. Ren G, Rudenko G, Ludtke SJ, Deisenhofer J, Chiu W, Pownall HJ. Proc Natl Acad Sci U. S. A. 107(3):1059-64 (2010).
15. Mechanism of LDL binding and release probed by structure-based mutagenesis of the LDL receptor. Huang S, Henry L, Ho YK, Pownall HJ, Rudenko G. J Lipid Res. 51(2):297-308 (2010). PubMed PMID:
14. Regulation of neurexin 1β tertiary structure and ligand binding through alternative splicing. Shen KC, Kuczynska DA, Wu IJ, Murray BH, Sheckler LR, Rudenko G. Structure 16(3):422-31 (2008).
Highlight: Featured article by Structure13. Dimerization and DNA-binding properties of the transcription factor DeltaFosB. Jorissen HJ, Ulery PG, Henry L, Gourneni S, Nestler EJ, Rudenko G. Biochemistry 46(28):8360-72 (2007).
12. Crystal structure of the second LNS/LG domain from neurexin 1alpha: Ca2+ binding and the effects of alternative splicing. Sheckler LR, Henry L, Sugita S, Südhof TC, Rudenko G. J Biol Chem. 281(32):22896-905 (2006).
11. Regulation of DeltaFosB stability by phosphorylation. Ulery PG, Rudenko G, Nestler EJ. J Neurosci. 26(19):5131-42. (2006).
10. 'MAD'ly phasing the extracellular domain of the LDL receptor: a medium-sized protein, large tungsten clusters and multiple non-isomorphous crystals. Rudenko G, Henry L, Vonrhein C, Bricogne G, Deisenhofer J. Acta Crystallogr D Biol Crystallogr. 59(Pt 11):1978-86 (2003).
9. The low-density lipoprotein receptor: ligands, debates and lore. Rudenko G, Deisenhofer J. Curr Opin Struct Biol. 13(6):683-9.(2003) Review.
8. Structure of the LDL receptor extracellular domain at endosomal pH. Rudenko G, Henry L, Henderson K, Ichtchenko K, Brown MS, Goldstein JL, Deisenhofer J. Science . 298(5602):2353-8 (2002).
Highlight: Comment in Science 298(5602):2337-9 (2002)7. LG/LNS domains: multiple functions - one business end? Rudenko G, Hohenester E, Muller YA. Trends Biochem Sci. 26(6):363-8 (2001) Review.
6. The structure of the ligand-binding domain of neurexin 1β: regulation of LNS domain function by alternative splicing. Rudenko G, Nguyen T, Chelliah Y, Südhof TC, Deisenhofer J. Cell. 99(1):93-101 (1999).
5. The atomic model of the human protective protein/cathepsin A suggests a structural basis for galactosialidosis. Rudenko G, Bonten E, Hol WG, d'Azzo A. Proc Natl Acad Sci U S A. 95(2):621-5. (1998).
4. Structure determination of the human protective protein: twofold averaging reveals the three-dimensional structure of a domain which was entirely absent in the initial model. Rudenko G, Bonten E, d'Azzo A, Hol WG. Acta Crystallogr D Biol Crystallogr. 52(Pt 5):923-36 (1996).
3. Three-dimensional structure of the human 'protective protein': structure of the precursor form suggests a complex activation mechanism. Rudenko G, Bonten E, d'Azzo A, Hol WG. Structure. 3(11):1249-59 (1995).
2. N-glycosylation and deletion mutants of the human MDR1 P-glycoprotein. Schinkel AH, Kemp S, Dollé M, Rudenko G, Wagenaar E. J Biol Chem. 268(10):7474-81 (1993).
1. In search of new lead compounds for trypanosomiasis drug design: a protein structure-based linked-fragment approach. Verlinde CL, Rudenko G, Hol WG. J Comput Aided Mol Des. 6(2):131-47 (1992).