Software
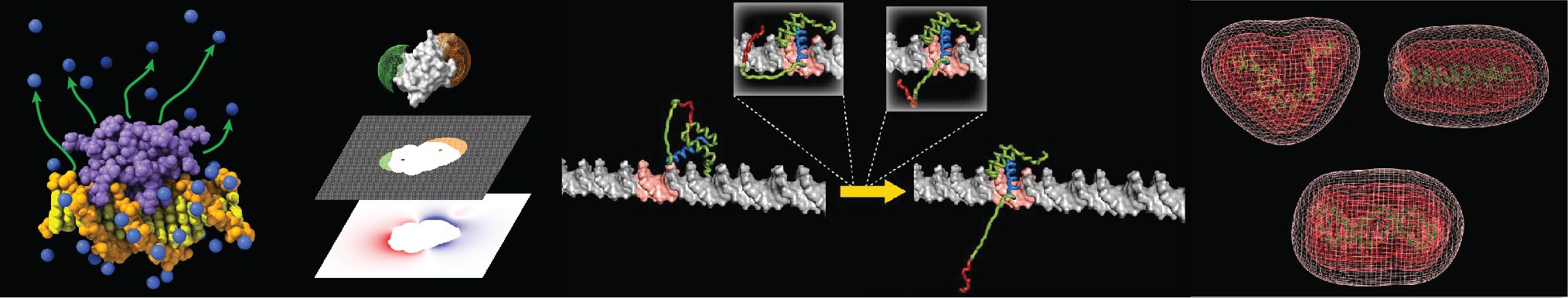
The following programs were developed by our group. Click each link for more information and references. If you would like to use them, please contact Junji Iwahara. Download instructions will be sent via email.
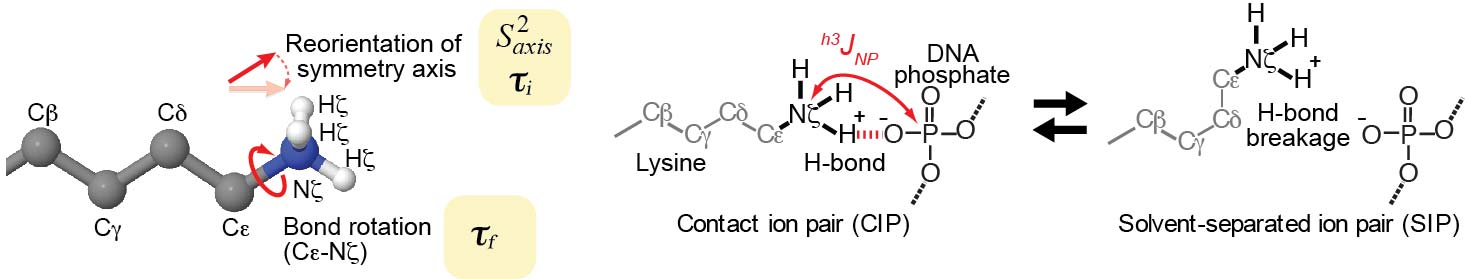
NMR pulse programs to study dynamics of lysine side-chain NH3+ groups
NMR pulse programs to study dynamics of lysine side-chain NH3+ groups
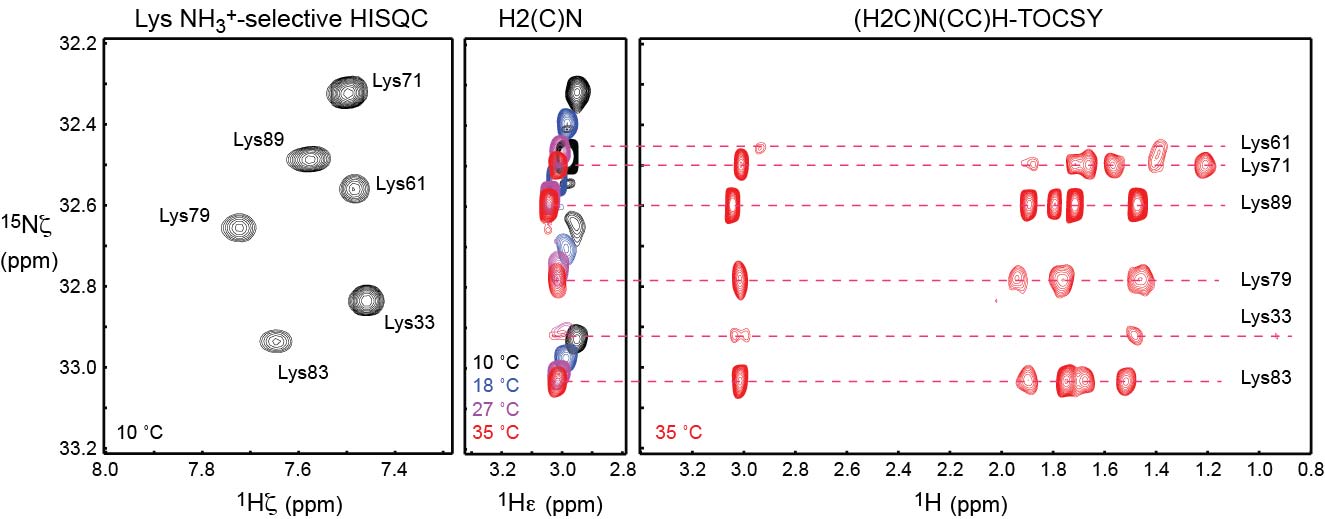
NMR pulse programs for triple resonance experiments to assign lysine NH3+ groups
NMR pulse programs for triple resonance experiments to assign lysine NH3+ groups
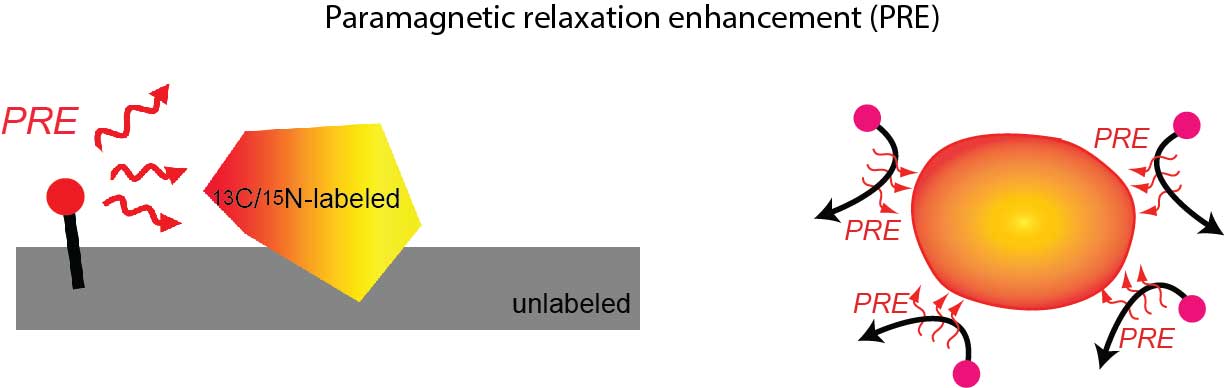

NMR pulse programs for 1H PRE rates Γ1 and Γ2 measurements
NMR pulse programs for 1H PRE rates Γ1 and Γ2 measurements
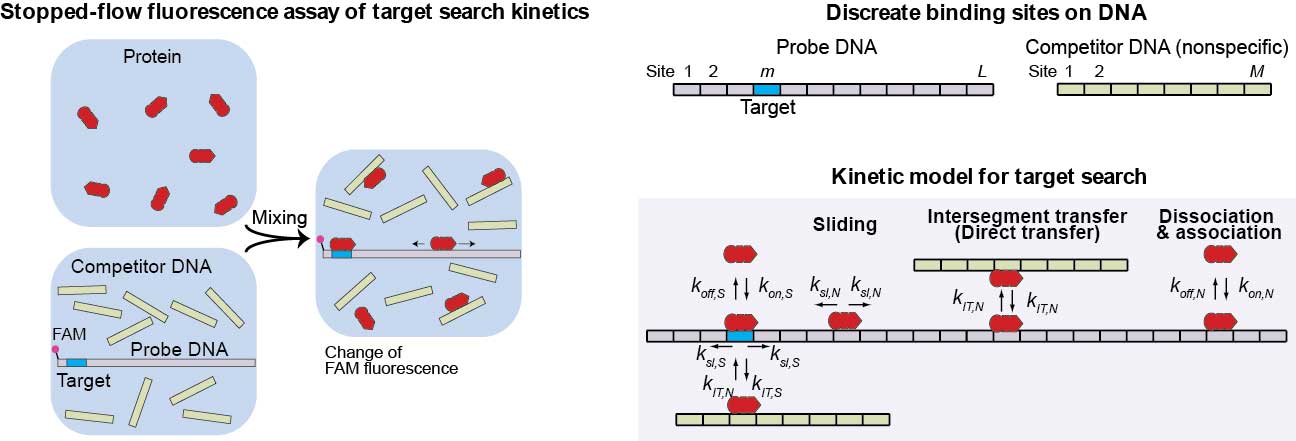
TDSK for kinetic analyses of protein translocation on DNA
TDSK for kinetic analyses of protein translocation on DNA