Andres F. Oberhauser, PhD 
Professor, Department of Neuroscience & Cell Biology
Tel: (409) 772-1309
Fax: (409) 747-2187
E-mail: afoberha@utmb.edu
Campus Location:5.212A Research Bldg 17
Mail
Route: 06203
Research
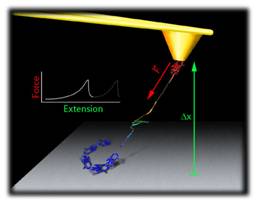
Single-molecule methods have emerged as powerful tools in life science research. These techniques allow the detection and manipulation of individual biological molecules and investigate, with unprecedented resolution, their conformations and dynamics at the nanoscale level. These techniques overcome the restrictions of traditional bulk biochemical studies by focusing on individuals of molecules. Our research focuses on the dynamics and mechanics of proteins using single-molecule manipulation techniques. Research Highlights: i) Mutations in Polycystin-1 cause polycystic kidney disease which is a common life-threatening genetic disease. We have discovered that Polycystin-1 has unique mechanical properties (Qian et al., J Biol Chem 2005; Xu et al., J. Biophysics 2013) and that pathogenic mutations can alter its nano-mechanics (Ma et al, J Biol Chem 2009, 2010); ii) We have found that titins are finely tuned to their micro-environment and that titin domains can fold under an applied force which hints for a previously uncharacterized folding-based spring mechanism (Bullard et al., PNAS 2006). We discovered that titin protein kinase domains have mechanical properties that are consistent with a function as effective force sensor (Greene et al, Biophys J. 2008); iii) Elastin and collagens are key structural component of the extracellular matrix.
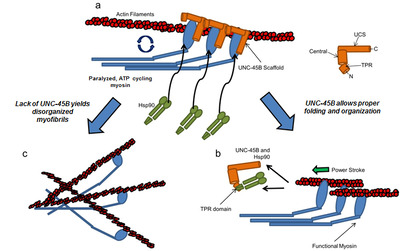
Despite its fundamental importance in tissue elasticity very little is known about the mechanical properties of native elastin and collagen fibers at the nano-molecular level. We have found that single tropoelastin molecules (a soluble precursor of elastin) can be stretched/relaxed hundreds of times, with no signs of hysteresis or molecular fatigue (Baldock et al., PNAS 2011). These mechanical properties make elastin an ideal molecular spring which is perfectly designed to undergo many stretch/relaxation cycles during the normal operation of different tissues (Holst et al., Nature Biotechnol. 2010); iv) Little is known about the molecular mechanisms that mediate myosin biogenesis into semi-crystalline arrays in muscle cells. We know that the molecular chaperones UNC-45 and Hsp90 are involved, but the actual mechanism has remained enigmatic. Using in vitro assays, we have now found surprising results regarding the interactions between UNC-45 and Hsp90 with myosin (Kaiser et al., Biophys J. 2012; Bujalowski et al., Biophys J. 2014; Nicholls et al., FEBS let 2014).
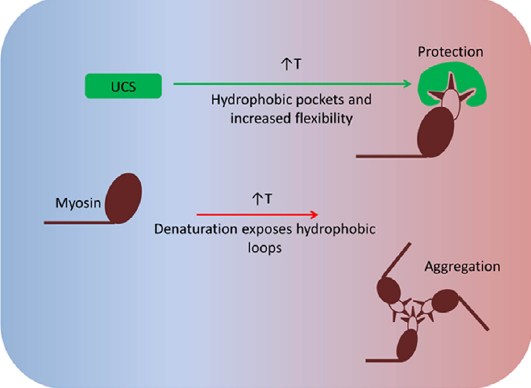
A keystone of our findings is a novel chaperone bound state of myosin that, to our knowledge, reconciles all previous genetic, biochemical and structural data pertaining to this system. v) The muscle-specific molecular chaperone UNC-45B is known to be involved in myosin folding and is trafficked to the sarcomeres A-band during thermal stress. We recently identified a temperature-dependent structural change in the UCS chaperone domain of UNC-45B that occur within a physiologically relevant heat-shock range. We show that distinct changes to the armadillo repeat protein topology result in exposure of hydrophobic patches, and increased flexibility of the molecule (Bujalowski et al; FEBS lett 2015). We suggest that these changes may function to suppress aggregation under stress by allowing binding to a wide variety of aggregation prone loops on its myosin client.
