 Thomas Smith, PhD
Thomas Smith, PhD
Professor, Department of Biochemistry & Molecular Biology
Tel: (409) 772-6028
E-mail: thosmith@utmb.edu
Campus Location: 5.104D Basic Science Bldg
Mail Route: 0645
Pubmed Publications | Lab Webpage
Research
The Smith Lab is currently working on the structures (cryo-EM and crystallographic) of a number of viruses, ABC transporters, carbohydrate recognition complexes, enzymes, and are using biochemical methods to protect crops with antifungal proteins.
Noroviruses
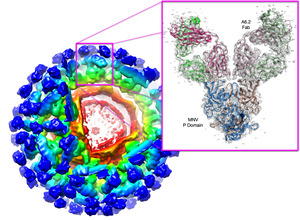 Noroviruses (family Caliciviridae) are the major cause of epidemic nonbacterial gastroenteritis in humans but the mechanism of antibody neutralization is unknown, and no structure of an infectious virion has been reported. Murine norovirus (MNV) is the only norovirus that can be grown in tissue culture, studied in an animal model, reverse engineered via an infectious clone, and to which neutralizing antibodies have been isolated. We have determined the cryo-electron microscopy structures of an MNV virion and the virion in complex with neutralizing Fab fragments and the atomic structure of the neutralizing antibody. As part of a larger DARPA project, we have been using a variety of methods to predict what kinds of mutations in the virus could lead to escape from antibody neutralization with the hope to extrapolate these findings to other virus systems that have more lethal pandemics. |
Plant Viruses
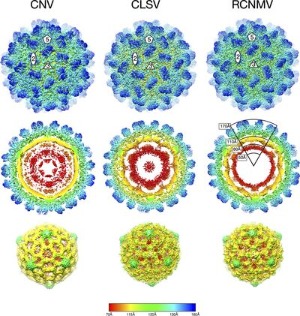 While many viruses are transmitted by various vectors (e.g. insects and fungi), little is know about the molecular interactions between the virus vectors (e.g. insects and fungi), little is know about the molecular interactions between the virus and the vector. To this end, we determined the crystal and cryo-EM structures of cucumber mosaic virus (CMV) and cucumber necrosis virus (CNV). CMV is a plant pathogenic virus of the family Bromoviridae that is readily transferred by aphids and survives on a wide variety of plants. We have localized the key element on the virus surface necessary for aphid transmission. CNV is transmitted in nature via zoospores of the Chytridiomycete fungus Olpidium bornovanus. We found using crystallography and cryo-EM that the virus is composed of several shells that seem to play a major role in vector transmission and we are currently examining the conformational changes associated with transmission. |
Glutamate Dehydrogenase and Insulin Disorders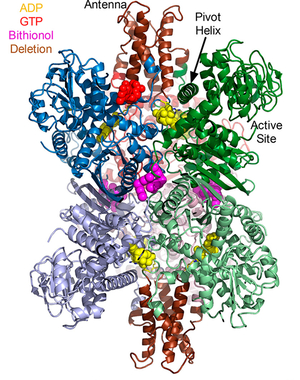 Mammalian glutamate dehydrogenase (GDH) is a mitochondrial enzyme that catalyzes the reversible oxidative deamination of L-glutamate to 2-oxoglutarate
using NAD(P)+ as coenzyme. The enzyme is a homohexamer that is tightly regulated by a large number of positive and negative allosteric effectors as well as by cooperative interactions between subunits. While the enzyme is found in all organisms, this
regulation is only found in the animal form. In collaboration with Charles Stanley's laboratory, we found that elimination of an inhibitory (GTP binding) site causes hyperinsulinism/hyperammonemia (HHI). Most interestingly, we found that compounds
from green tea are effective in controlling HHI in transgenic mice expressing the defective human form of GDH.
Mammalian glutamate dehydrogenase (GDH) is a mitochondrial enzyme that catalyzes the reversible oxidative deamination of L-glutamate to 2-oxoglutarate
using NAD(P)+ as coenzyme. The enzyme is a homohexamer that is tightly regulated by a large number of positive and negative allosteric effectors as well as by cooperative interactions between subunits. While the enzyme is found in all organisms, this
regulation is only found in the animal form. In collaboration with Charles Stanley's laboratory, we found that elimination of an inhibitory (GTP binding) site causes hyperinsulinism/hyperammonemia (HHI). Most interestingly, we found that compounds
from green tea are effective in controlling HHI in transgenic mice expressing the defective human form of GDH.
Human Rhinovirus
Human rhinovirus is one of the major causes of the common cold. There are over 100 serotypes of this virus, making it unlikely that there will ever be a traditional vaccine for the common cold using conventional
vaccine methods. The Smith lab has examined the neutralization and dynamics of this virus to better understand how antibodies block infection and how to make more efficacious and novel vaccines.
Antifungal Proteins
We have determined the structures of antifungal proteins, KP4 and KP6, that are expressed by viruses that persistently infect Ustilago maydis. While KP4 acts by blocking voltage gated calcium channels in the target
fungi, the mode of action of KP6 is unclear other than it lyses the target cell. Since both KP4 and KP6 affect other Ustilago maydis, it may be possible to leaverage these proteins to treat human pathogens as well.
ABC Transporters
We have determined the structures of a number of ion binding domains from the ABC transporters of Synechocystis 6803. In our first study, determined the structure of the periplasmic domain of a zinc transporter,
ZnuA investigated structural aspects of the protein that may be involved in import regulation. Bioavailable iron is a limiting nutrient for primary production in large areas of the oceans. We determined the atomic structure of the ferric binding protein,
FutA, and examined the conformational changes in the protein as it binds metal. Cyanobacteria are the most abundant microorganisms in aquatic environments and play a key role in the global carbon cycle. To better understand how it concentrates CO2, we determined the atomic structure of the bicarbonate binding protein, CmpA. Having already determined the structure of a nitrate transporter, NrtA, we found clues to how these proteins are remarkably selective and how they may have 'cross talk' regulation.
Carbohydrate binding and processing proteins
Our gut microbiota recognize and process undigestible carbohydrates in the human gut. To better understand how they are able to process such a diverse collection of polysaccharides, we
determined the structures of a number of bacterial proteins that bind (SusD and Bt1043) and process (SusG) these carbohydrates.
