 B. Montgomery Pettitt, PhD
B. Montgomery Pettitt, PhD
Director, Sealy Center for Structural Biology and Molecular Biophysics
Robert A. Welch Distinguished Chair in Chemistry, Department of Pharmacology & Toxicology
Professor, Departments of Pharmacology & Toxicology, and Biochemistry & Molecular Biology
Tel: (409) 772-0723
Fax: (409) 772-0725
E-mail: mpettitt@utmb.edu
Campus Location: 5.404A Research Bldg 6
Mail
Route: 0304
Pubmed Publications | Lab Webpage
Research
The research in my laboratory involves work in the areas of : (1) Biochemistry, (2) Chemical Physics/Physical Chemistry and (3) Computer Science. In particular, I am interested in
Computational theory of exotic statistical ensembles and sampling methods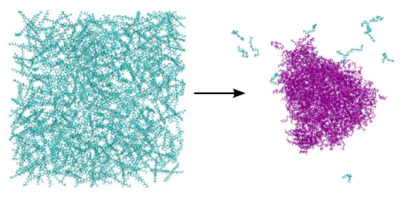 In particular the use of chemical potential to calculate phase related behavior requires sampling tricks. Application to systems at constant activity to explore phase transitions in saline solution and protein folding in multicomponent systems is of interest.
In particular the use of chemical potential to calculate phase related behavior requires sampling tricks. Application to systems at constant activity to explore phase transitions in saline solution and protein folding in multicomponent systems is of interest.
Forces and structures governing thermodynamics and kinetics in liquid solutions especially aqueous systems
Most difficult is the question of how multicomponent systems including cosolvents and ions affect the structure of proteins and nucleic acids
in solution. Given correlations and statistical thermodynamics the relations to experimental observables on the effects ions and osmolytes have on biomacromolecules in solution should then be understandable. At the technical level we are working
on activity models and diagramatic expansion.
DNA confinement
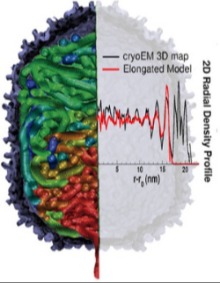 Bacteriophages, found in bacteria-rich locations like rivers and soil, are nature's machinery for viral infection of bacteria. Their genetic material, DNA or RNA, single- or double-stranded, are carried in protein-based capsids and released into the bacteria. Understanding the biophysical basis of the biological process which transfers a viral genome to infect a cell is important to the cellular machinery and many disease related fields. DNA, a charged elastic polymer, undergoes over 250-fold compaction when packed into a capsid overcoming an unfavorable thermodynamic barrier by using ATP. How DNA overcomes the unfavorable thermodynamic barrier to enter and pack inside a capsid depends on the interplay of many different intermolecular interactions. Combined with experimental data, coarse-grained models and multi-scale techniques are being employed to model the structure and, consequently, the thermodynamics of DNA confined by surfaces. |
Effects of anisotropic environments on DNA and Proteins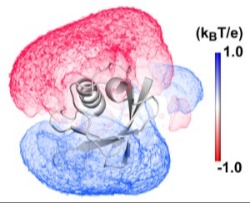 The electric fields, solvent gradients and density waves near surfaces have profound effects on conformation and binding. Both theoretical and simulation methods are being developed to address these problems.
The electric fields, solvent gradients and density waves near surfaces have profound effects on conformation and binding. Both theoretical and simulation methods are being developed to address these problems.
Theory and computational methods to investigate solution systems with couplings and correlations at many disparate length and time scales. There are many problems for which atomic correlations do not provide a direct link to macroscopic properties. Connecting meso scale averaging procedures to the atomic and macro levels via multiscale methods is important for biological/materials applications.
