 Stanley J. Watowich, PhD
Stanley J. Watowich, PhD
Professor, Department of Biochemistry & Molecular Biology
Tel: (409) 747-4749
Fax: (409) 747-4745
E-mail: watowich@bloch.utmb.edu
Campus Location:623 Basic Science
Bldg
Mail Route: 0645
Research
Students and post-docs in my laboratory are (1) studying the structure and assembly of RNA enveloped viruses, (2) working to discover of novel drugs and drug targets to combat infectious diseases, and (3) constructing a quantitative mechanistic model to completely describe receptor tyrosine kinase activation and signaling. Unifying these projects is a desire to understand the structural and biophysical principles that regulate complex biomolecular interactions, and to predictably modulate those interactions responsible for disease states. Progress towards these ends is achieved from combining structural, biochemical, biophysical, genomic, proteomic, and molecular biology approaches.
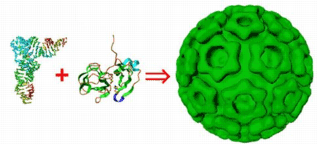 (1) We have determined the structure of the capsid protein of Venezuelan equine encephalitis virus (family: Togaviridae) and are characterizing the structural transitions that occur during the assembly and disassembly of this enveloped
ssRNA virus. The kinetics of virus assembly are being investigated through the use of a reconstituted in vitro assembly system. In addition, we are studying the structure, function, and assembly of the core and envelope proteins from hepatitis C virus.
This integrated information will be used to efficiently screen combinatorial libraries as part of our structure-based drug design program to develop agents to inhibit viral assembly and cell-surface attachment.
(1) We have determined the structure of the capsid protein of Venezuelan equine encephalitis virus (family: Togaviridae) and are characterizing the structural transitions that occur during the assembly and disassembly of this enveloped
ssRNA virus. The kinetics of virus assembly are being investigated through the use of a reconstituted in vitro assembly system. In addition, we are studying the structure, function, and assembly of the core and envelope proteins from hepatitis C virus.
This integrated information will be used to efficiently screen combinatorial libraries as part of our structure-based drug design program to develop agents to inhibit viral assembly and cell-surface attachment.
 (2) Current approaches to developing antipathogen therapeutics are extremely costly and time-consuming, but rarely successful. To address this problem, we have implemented a Functional Phenotype screening technology to rapidly identify
novel drug leads and drug targets to combat infectious pathogens. Using a variety of proprietary cell-based combinatorial libraries, we have successfully generated independent cell lines that resist anthrax toxin and all tested alphaviruses. Detailed
differential genomic, proteomic and transgenic analysis of these discretely modified cells is used to identify and validate critical antipathogen targets and pathways. When fully deployed, this technology, together with our new BSL-4 biocontainment
laboratories, can be mobilized to rapidly identify validated drug leads and targets for most BWT and emerging/reemerging pathogens.
(2) Current approaches to developing antipathogen therapeutics are extremely costly and time-consuming, but rarely successful. To address this problem, we have implemented a Functional Phenotype screening technology to rapidly identify
novel drug leads and drug targets to combat infectious pathogens. Using a variety of proprietary cell-based combinatorial libraries, we have successfully generated independent cell lines that resist anthrax toxin and all tested alphaviruses. Detailed
differential genomic, proteomic and transgenic analysis of these discretely modified cells is used to identify and validate critical antipathogen targets and pathways. When fully deployed, this technology, together with our new BSL-4 biocontainment
laboratories, can be mobilized to rapidly identify validated drug leads and targets for most BWT and emerging/reemerging pathogens.  In addition to our holistic Functional Phenotype filtering approach, we are extending traditional computer-based drug discovery approaches to identify compounds with antiviral activity in cell culture and animal models. We use a synergistic combination
of high-throughput in vitro and in silico screening, X-ray crystallography, homology modeling, and novel bioavailability filters, to identify compounds that effectively disrupt alphavirus (e.g., Venezuelan equine encephalitis virus) and flavivirus
(e.g., dengue and West Nile viruses) replication in vivo.
In addition to our holistic Functional Phenotype filtering approach, we are extending traditional computer-based drug discovery approaches to identify compounds with antiviral activity in cell culture and animal models. We use a synergistic combination
of high-throughput in vitro and in silico screening, X-ray crystallography, homology modeling, and novel bioavailability filters, to identify compounds that effectively disrupt alphavirus (e.g., Venezuelan equine encephalitis virus) and flavivirus
(e.g., dengue and West Nile viruses) replication in vivo.
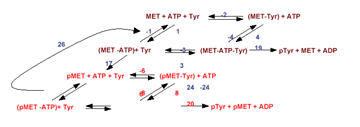 (3) We are investigating the conformational and thermodynamic changes responsible for the activation and signaling of cell-surface receptor tyrosine kinases (RTKs). For these studies we have developed soluble molecules that mimic
the activated c-MET RTK, a proto-oncoprotein implicated in tumorogenesis and metastasis. This reconstituted system is amenable to X-ray crystallographic and biochemical studies. Significantly, we have measured the thermodynamic and kinetic changes
that accompany receptor oligomerization. This biochemical data has been incorporated into a detailed quantitative model that explains how oligomerization facilitates RTK activation and signaling. In addition, our soluble c-MET RTK is a target for
in vitro and computer-based drug discovery studies that are looking to find inhibitors of receptor phosphorylation, and thus may serve as anti-metastasis agents.
(3) We are investigating the conformational and thermodynamic changes responsible for the activation and signaling of cell-surface receptor tyrosine kinases (RTKs). For these studies we have developed soluble molecules that mimic
the activated c-MET RTK, a proto-oncoprotein implicated in tumorogenesis and metastasis. This reconstituted system is amenable to X-ray crystallographic and biochemical studies. Significantly, we have measured the thermodynamic and kinetic changes
that accompany receptor oligomerization. This biochemical data has been incorporated into a detailed quantitative model that explains how oligomerization facilitates RTK activation and signaling. In addition, our soluble c-MET RTK is a target for
in vitro and computer-based drug discovery studies that are looking to find inhibitors of receptor phosphorylation, and thus may serve as anti-metastasis agents.
